- Number 413 |
- May 12, 2014
Study reveals genetic diversity in Prochlorococcus
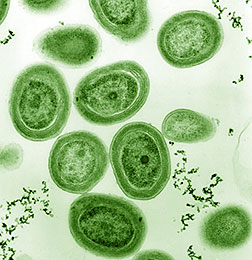
Electron micrograph of a cultured strain of
Prochlorococcus, MED4.
(N. Watson and L. Thompson, MIT)
A million cells of Prochlorococcus can be found in a single liter of water, and this organism is thought to be responsible for providing about 20% of the oxygen produced by the planet each year. The cyanobacterium can be classified into several distinct major ecotypes – characterized by variable factors such as seasonal, depth, and geographic patterns.
However, even within these ecotypes,Prochlorococcus cells still display a wide range of genomic diversity. In the April 25, 2014 issue of Science, MIT marine microbiologist Sallie Chisholm, a longtime collaborator of DOE Joint Genome Institute (DOE JGI), led a team that applied single-cell genomics to these cyanobacteria collected from the same environment at three separate times of the year.
The researchers sequenced and assembled Prochlorococcus genomes from single cells collected at the Bermuda-Atlantic Time-series Study site in the northwestern Sargasso Sea between November 2008 and April 2009. The team wondered if the Prochlorococcus subpopulations within the clusters were genomically distinct. The answer turned out to be yes, and each of these so-called “genomic backbones” is comprised of highly conserved core gene alleles, or alternative forms, and a smaller distinct set of flexible genes. If each of the cyanobacterial backbone subpopulations were counted as distinct species, the team noted, then Prochlorococcus consists of thousands of species.
The vast diversity of these abundant Prochlorococcus subpopulations was validated with help from the DOE JGI’s Rex Malmstrom. “My contribution was to provide the essential control data,” he explained. “We could do this based on our extensive R&D work on single cell genomics.” Read more at http://jgi.doe.gov/discovering-diversity-one-cell-time/.[Massie Ballon, 925.927.2541,
mlballon@lbl.gov]
